This project is archived and no longer maintained.
BayesMix
| Overview | |
| Description | A tool for performing model-based inference for differential gene expression, using a non-parametric Bayesian mixture probability model for the distribution of gene intensities under different conditions |
| Development Information | |
| Language | R and C |
| Current version | 0.8.8 |
| Platforms | Windows and Linux |
| Status | Inactive |
| Last updated | 15th August 2003 |
BayesMix
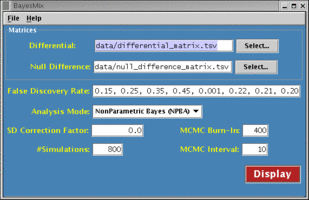
BayesMix is a piece of computer software, a suite of R functions and C routines with a graphical user interface, which can be used as a tool for performing model-based inference for differential gene expression, using a non-parametric Bayesian mixture probability model for the distribution of gene intensities under different conditions. For comparison purposes, Efron’s empirical Bayes method (JASA, 2001), is also implemented. The package can be employed for any general research problem that involves massive multiple comparisons with continuous outcomes.
The package produces numerous plots and tables from which the user can:
- estimate the probability of differential gene expression for each gene with full posterior uncertainty
- estimate the total number of significantly differential genes for different levels of false discovery rates, with automatic Bayesian multiplicity adjustment
- easily perform joint inferences on multiple genes
Download
The current Un*x-based version is 0.8.8, available as of 2003/08/15. Here are some links to download various binary packages. The source package is not publicly available at this time since it’s still being written. The following are the supported platforms.
- Sun Solaris 2.8
- Compaq Tru64 4.0f
- RedHat Linux 7.1 (IA32)
- SuSE Linux 8.0 (IA32)
- Mac OS X 10.2 (Jaguar)
- Windows 98, 2000, XP
System Requirements
Documentation
Credits
- Kim-Anh Do, Ph.D.
- Bradley Broom, Ph.D.
- Peter Müller, Ph.D.
- Sijin Wen
- Paul Roebuck
- Spyros Tsavachidis
References
Kim-Anh Do, Peter Müller, Feng Tang (2003).
Bayesian Mixture Model for Differential Gene Expression.
Technical Report, Department of Biostatistics, University of Texas/MD Anderson Cancer Center
Bradley Efron, Robert Tibshirani, John D. Storey, Virginia Tusher (2001).
Empirical Bayes Analysis of a Microarray Experiment.
Journal of the American Statistical Association, Volume 96, Number 456, Pages 1151-116