Available Software
Below are software and services provided by the Department of Bioinformatics and Computational Biology. These tools are copyrighted by the University of Texas MD Anderson Cancer Center and by the individual employees of the cancer center who helped develop them. They are freely available for personal use in research projects; however, anyone wishing to use them or modify them for use in a commercial project should contact the Office of Technology Commercialization.Featured Software

All Software
-
BM-Map
A Powerful Bioinformatics Software Tool for Refining Next-Generation Sequencing (NGS) Read Mapping
-
BreakDown
BreakDown genotyping SVs in heterogeneous tumor sample
-
BreakFusion
A computational pipeline for identifying gene fusions from RNA-seq data.
-
BreakTrans
BreakTrans maps gene fusions to genomic architecture in cancer cells
-
CanDrA
CanDrA is a computer program that predicts cancer-type specific driver missense mutations
-
CliPP
Clonal structure identification through penalizing pairwise differences
-
DeMixT
Cell type-specific deconvolution of heterogeneous tumor samples with two or three components using expression data from RNAseq or microarray platforms
-
drugBase
drugBase provides a RESTful interface to a database of drug-target interactions.
-
DyCE
DyCE is a server for enabling remote users to access advanced computational modeling and analysis tools and view the results of the analysis directly in their browser.
-
ESTIMATE
A tool for predicting tumor purity, and the presence of infiltrating stromal/immune cells in tumor tissues using gene expression data.
-
Famdenovo
Estimate the probability of a germline mutation to be de novo using family history data.
-
FamSeq
FamSeq is a computational tool for calculating probability of variants in family-based sequencing data.
-
FASMIC
FASMIC is a comprehensive database of experimental-evidence-based functional impact of somatic mutations in cancer.
-
GeneClust
GeneClust is a tool used for exploratory analysis of gene expression microarray data. Available implementations include S-plus and R packages for installation on your machine, and a Javascript version for in-browser analysis of smaller datasets.
-
geneSmash
geneSmash is a mash-up of various sources of information about human genes including the NCBI Entrez gene FTP site, UCSC Genome Browser, miRBase and human gene expression array annotation extracted from Manufacturers’ websites. Use geneSmash
-
IdeogramViewer
An interactive tool for viewing the location and annotation of genes on a human chromosome.
-
LFSPRO
TP53 mutation is a main cause of Li-Fraumeni syndrome. This package is designed to estimate the probability that the counselee is a TP53 mutation carrier on the basis of his/her family cancer history. We also included LFS classic and Chompret criteria in the package.
-
MBatch
R package to help assess and correct for batch effects.
-
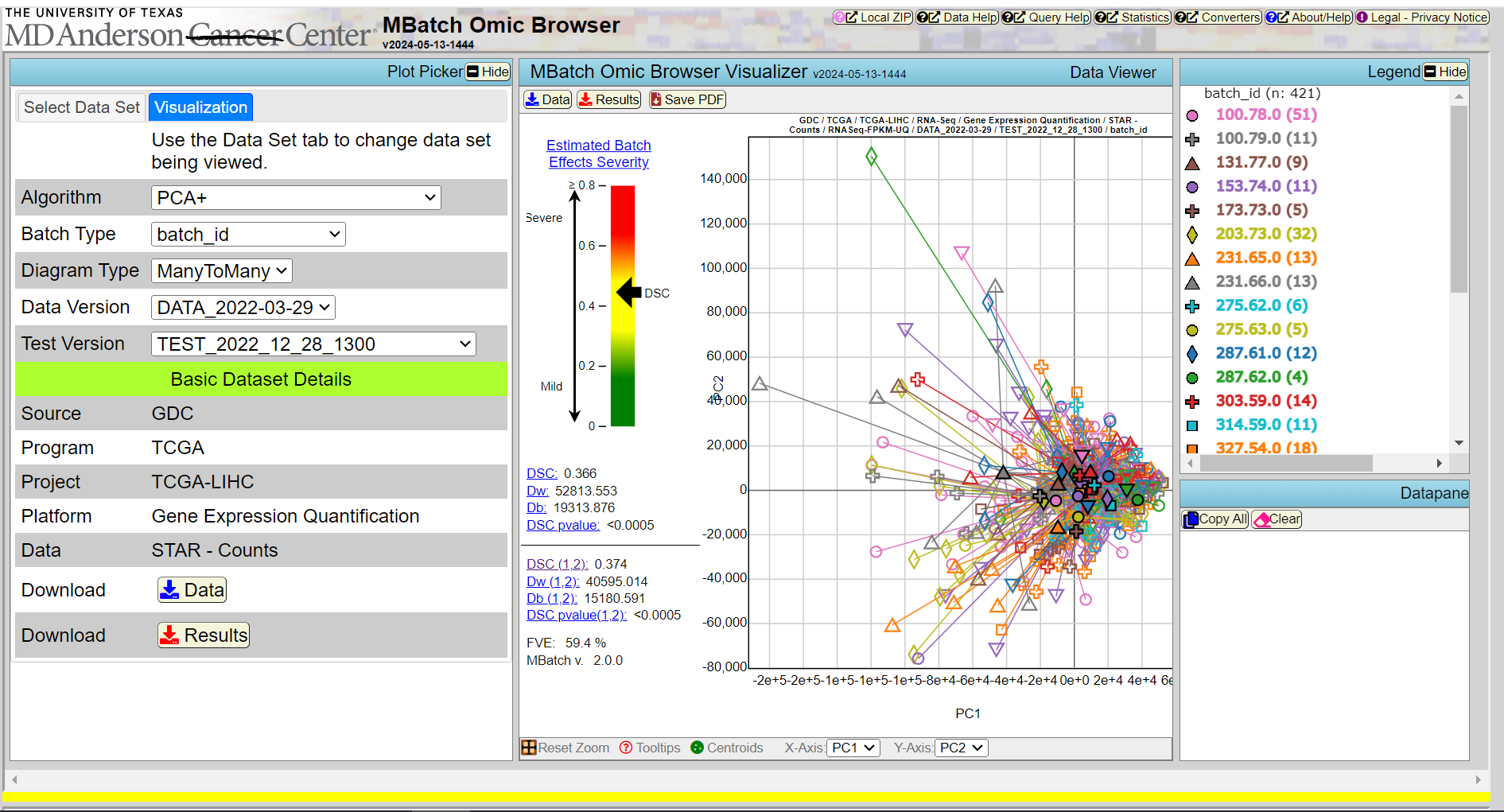
MBatch Omic Browser
MBatch Omic Viewer is a web-based tool for standardization and visualization of omics data.
-
MD Anderson Cell Lines Project (MCLP)
MCLP provides a comprehensive resource for accessing, visualizing, and analyzing functional proteomics of cancer cell lines.
-
MetaBatch Omic Browser
MetaBatch Omic Browser provides a graphical and statistical analysis of Metabolomics Workbench data sets performed using the MBatch R package with extensions for metabolomics data.
-
MuSE
Somatic point mutation caller for tumor-normal paired samples in next-generation sequencing data.
-
Next-Generation (Clustered) Heat Maps (NG-CHM)
Zoomable (clustered) heat maps with links to statistical information, databases, and other related analyses. This is Version 2 of the software, featuring a faster, more dynamic interface and a tool for building NG-CHMs within the Galaxy bioinformatics platform.
-
Pathways Web
An Open-Use Integrated API of Pathways, Genes, Directional Gene Interactions, and the Gene Ontology with Data Versioning for Provenance.
-
RDriver
Associating multi-omic data for accurate prediction of driver mutations in cancer
-
Ribo-TISH
Ribo-seq data-driven Translation Initiation Sites Hunter
-
RPPA SPACE
R package for analsys of RPPA data
-
SAMMI
SAMMI (Semi-Automated Metabolic Map Illustrator) is a web-based tool for the visualization of metabolic networks and related data.
-
Sample Sizes
A tool to compute the number of samples needed to detect expression changes for a microarray experiment. (Use Sample Sizes)
-
Script of Scripts
SoS Polyglot Notebook and SoS Workflow Engine, a multi-language environment for both interactive data analysis and batch data processing.
-
SpliceSeq
A tool for investigating alternative mRNA splicing in next generation mRNA sequence data
-
SuperCurve
SuperCurve is a stand-alone package, bundled with OOMPA, that provides tools for the analysis of reverse phase protein arrays.
-
SurvNet
This service provides a bioinformatic web app for identifying network-based biomarkers that most correlate with patient survival data. Use SurvNet.
-
SWAKK
This service provides a bioinformatic web app for positive selection in proteins using a sliding window substitution rate analysis. Use SWAKK.
-
TCGA Batch Effects
The Cancer Genomic Atlas (TCGA) project studies different types of cancer by obtaining datasets from different tissue source sites, using different sequencing centers and technologies. Data sets can sometimes be biased depending on the batch from which it came. The TCGA Batch Effects Tool provides pre-computed graphical annotations of different TCGA data sets that allows users to screen for batch biases in the dataset.
-
TCGA SpliceSeq
TCGA SpliceSeq presents an analysis of alternative mRNA splicing in thousands of TCGA cancer samples across 33 different tumor types. Percent splice in values for all possible splice events of all genes can be queried, graphically displayed, and downloaded.
-
The Atlas of non-coding RNA in Cancer (TANRIC)
An open-access resource for interactive exploration of lncRNAs in cancer.
-
The Cancer Mitochondrial Atlas (TCMA)
A comprehensive resource for accessing, visualizing, and analyzing cancer mitochondrial genomic data.
-
The Cancer Proteome Atlas (TCPA)
A comprehensive resource for accessing, visualizing, and analyzing cancer functional proteomics.
-
TransVar
TransVar is a multi-way annotator for genetic elements and genetic variations.